Global Health Sentiment Similarity (GDELT + PySpark)
Large-scale analysis of global health-news sentiment using GDELT, distributed MapReduce (PySpark), and vector-based NLP similarity methods.
Research question
How similar is the sentiment of health-related news coverage between New Zealand and other countries over time, and how can that similarity be computed efficiently over a large, distributed news dataset?
The emphasis is both analytical (sentiment similarity) and computational: designing a pipeline that scales to hundreds of country–month datasets using parallel processing.
Big Data source
This project uses the GDELT 2.0 global event database, which continuously aggregates and annotates news articles from across the world. I queried GDELT’s document API to retrieve health-related sentiment tone charts for multiple countries over a multi-year period.
Scale characteristics
- 222 CSV files (6 countries × 37 months)
- Each file aggregates thousands of news articles into tone-distribution buckets
- Data volume and structure motivated a distributed processing approach
While the per-file CSVs are modest in size, the overall workload reflects a common Big Data pattern: many independent data partitions requiring identical transformations.
Method: distributed sentiment aggregation
I implemented the analysis using PySpark to emphasize distributed computation and parallelisation. Each country–month dataset was processed using a MapReduce-style aggregation to compute an average sentiment score from tone buckets.
Parallel MapReduce design
- Map step: emit
(count)and(tone × count)pairs per bucket - Reduce step: aggregate sums across partitions
- Final step: compute weighted average sentiment per month
- Executed using Spark RDDs to allow parallel execution across nodes
NLP-inspired similarity
- Each country represented as a sentiment time-series vector
- Vectors compared using cosine similarity
- Discrete tone scaling applied to stabilize similarity metrics
Although this is not transformer-based NLP, the approach reflects a classic NLP workflow: transforming unstructured text into numerical representations and comparing documents (here, countries over time) in vector space.
Results
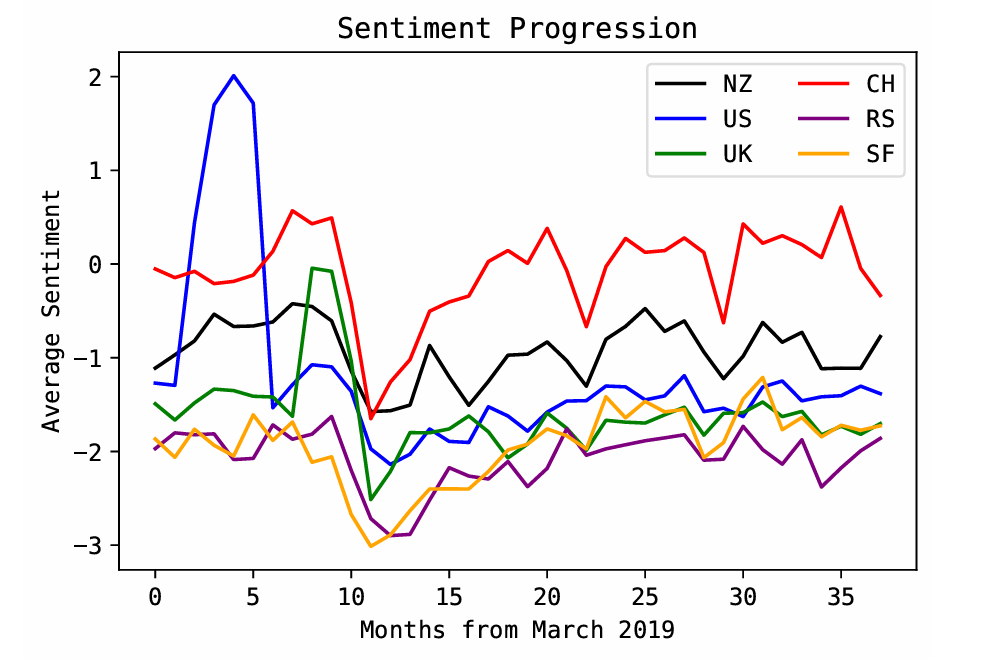
The sentiment progression was highly similar across all countries studied. New Zealand’s closest match was China (S ≈ 0.983) and the least similar was the United States (S ≈ 0.916), but all similarities remained high.
Key takeaway
The alignment of peaks/troughs across countries suggests a broadly consistent global media tone around health events during the COVID-era time window.
Parallelisation & performance analysis
A core goal of this project was to evaluate the effectiveness of parallelisation for this workload. I ran the pipeline across multiple Spark configurations to measure scaling behavior.
- Tested execution on 1, 2, 8, and 16 nodes
- Observed limited or negative speedup with increased parallelism
- Identified overhead from repeated small-RDD creation and forced sequential steps
Engineering insight
This result is itself instructive: parallel frameworks do not guarantee speedup. Effective Big Data systems require batching, minimizing driver-side logic, and structuring transformations to amortize scheduling and I/O costs.
What I’d improve next
Engineering
- Reduce small-RDD overhead by batching CSV ingestion
- Minimize Python-side loops; push aggregation into Spark transformations
- Cache/persist where appropriate; profile stages explicitly
Analysis
- Broaden country set and extend timeline beyond COVID-era
- Compare multiple keyword filters (e.g., “vaccine”, “pandemic”, “hospital”)
- Add uncertainty estimates (bootstrap months / resample sources)
Beyond the specific sentiment results, this project demonstrates end-to-end data science systems thinking: working with a global-scale dataset, designing parallel MapReduce-style aggregations in PySpark, applying NLP-inspired vector representations, and critically evaluating when distributed computation meaningfully improves performance — and when overhead dominates.